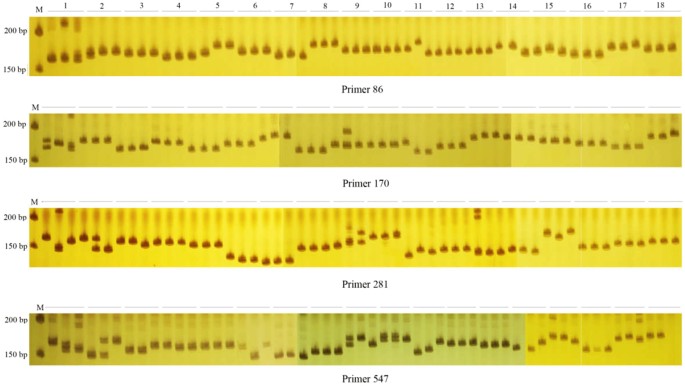
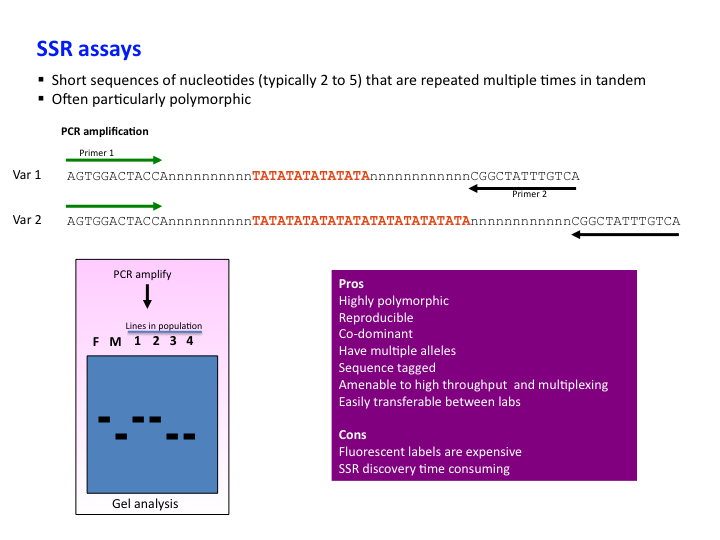
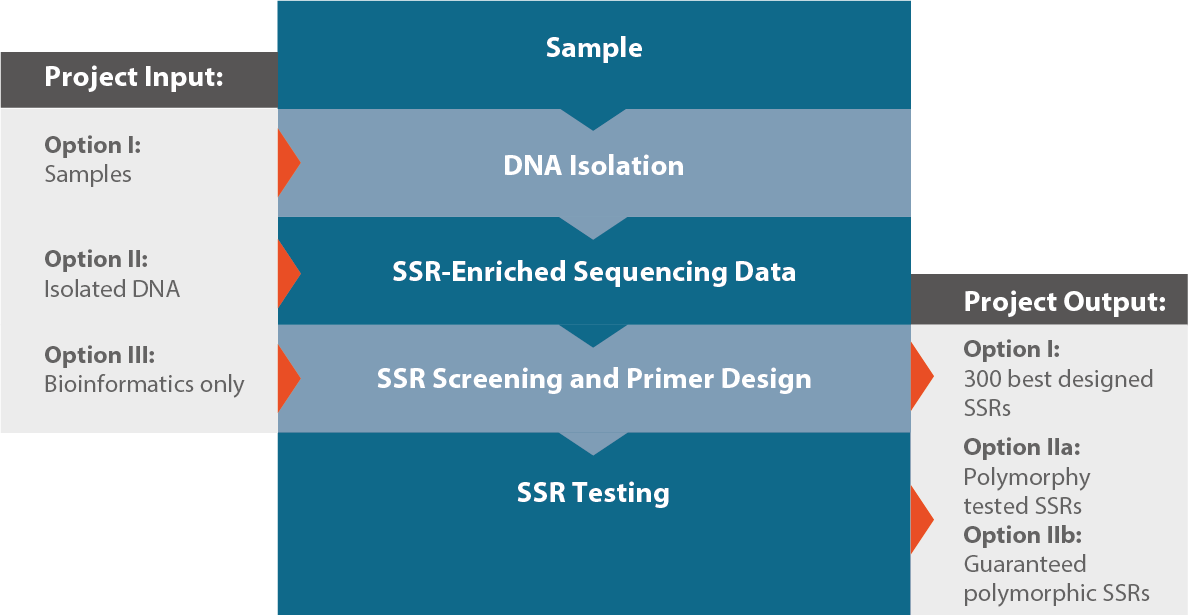
Development of unigene-derived SSR markers in cowpea (Vigna unguiculata) and their transferability to other Vigna species
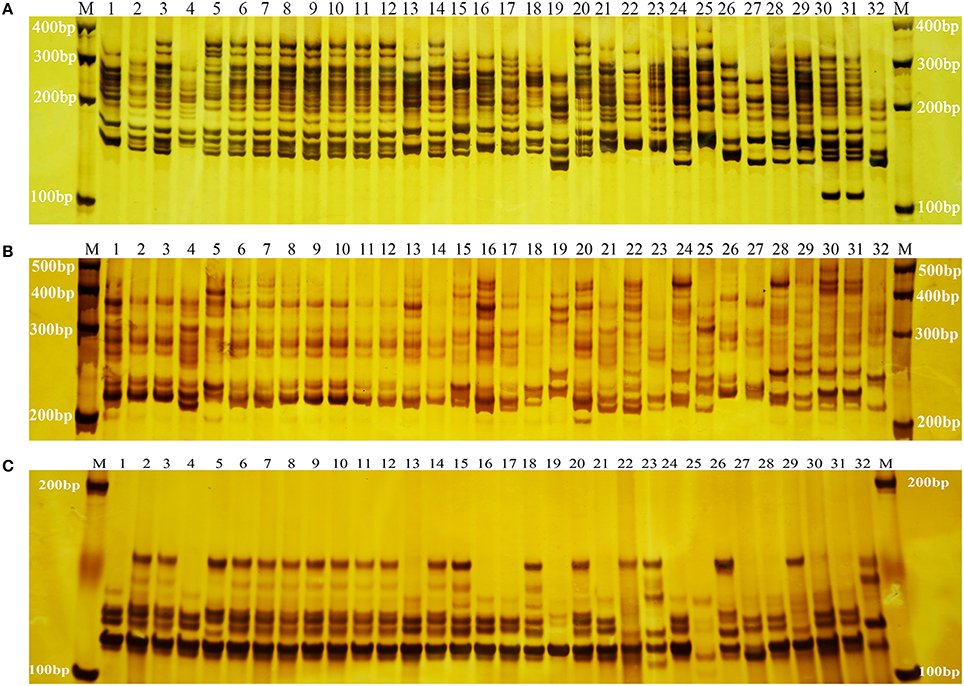
Frontiers | Development of SSR Markers and Assessment of Genetic Diversity in Medicinal Chrysanthemum morifolium Cultivars | Genetics

Comprehensive functional analysis and mapping of SSR markers in the chickpea genome (Cicer arietinum L.) - ScienceDirect
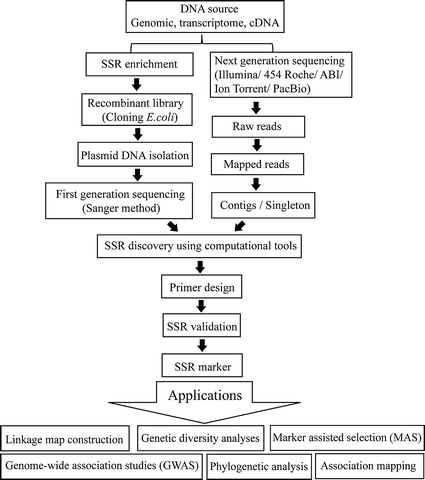
Simple sequence repeats (SSRs) markers in fish genomic research and their acceleration via next-generation sequencing and computational approaches | SpringerLink

Development of SSR markers by next-generation sequencing of Korean landraces of chamoe (Cucumis melo var. makuwa) | SpringerLink
![Plants | Free Full-Text | Analysis of Genetic Diversity and Population Structure of Pigeonpea [Cajanus cajan (L.) Millsp] Accessions Using SSR Markers Plants | Free Full-Text | Analysis of Genetic Diversity and Population Structure of Pigeonpea [Cajanus cajan (L.) Millsp] Accessions Using SSR Markers](https://www.mdpi.com/plants/plants-09-01643/article_deploy/html/images/plants-09-01643-g001.png)
Plants | Free Full-Text | Analysis of Genetic Diversity and Population Structure of Pigeonpea [Cajanus cajan (L.) Millsp] Accessions Using SSR Markers
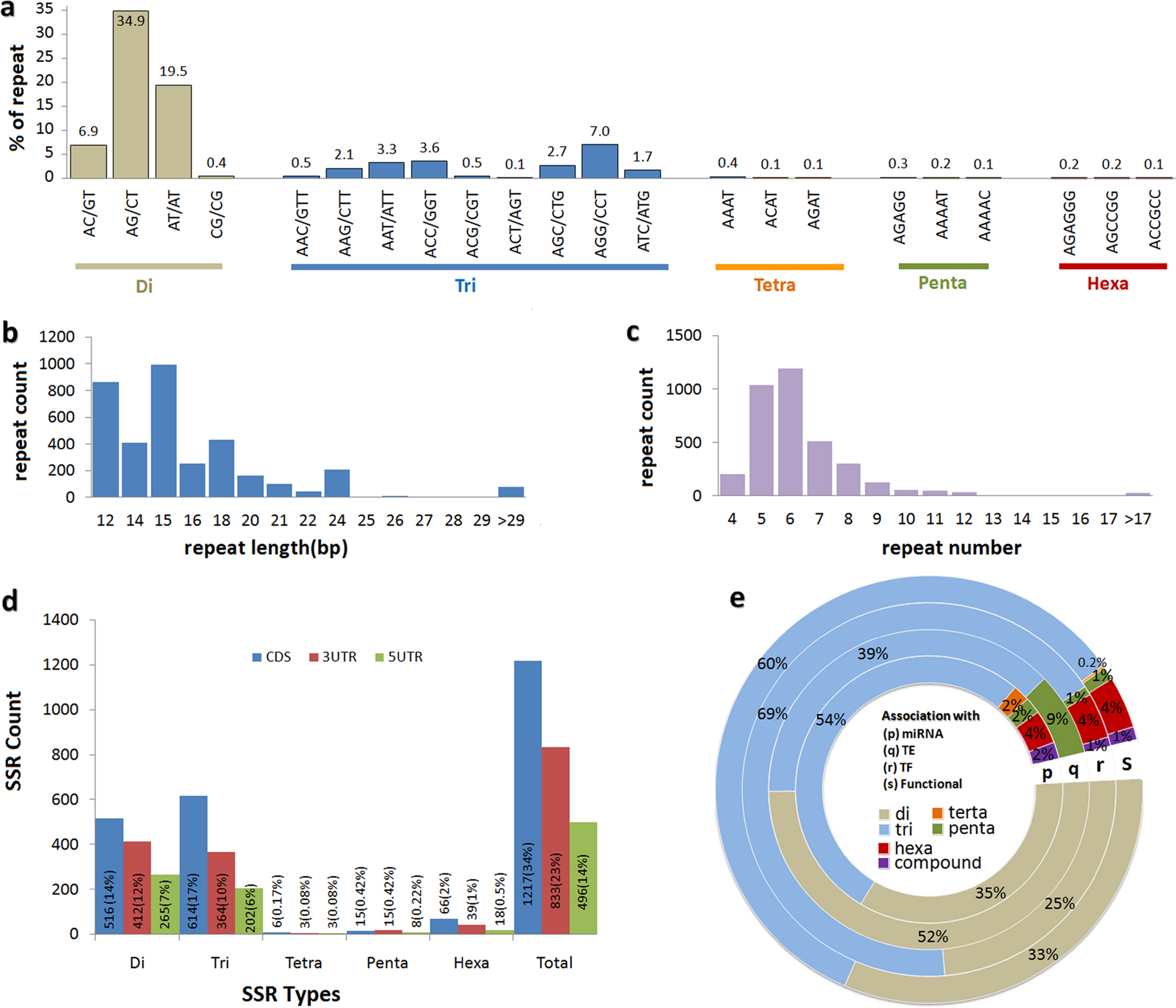
Transcriptome wide SSR discovery cross-taxa transferability and development of marker database for studying genetic diversity population structure of Lilium species | Scientific Reports

Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports
Genetic diversity of Venturia inaequalis isolates (Apple scab) in China and U.K. determined by SSR markers | PLOS ONE
Development of novel EST-SSR markers for ploidy identification based on de novo transcriptome assembly for Misgurnus anguillicaudatus | PLOS ONE

A general protocol for developing SSR markers with a SSR-enrichment step. | Download Scientific Diagram

SciELO - Brasil - Microsatellite markers: what they mean and why they are so useful Microsatellite markers: what they mean and why they are so useful
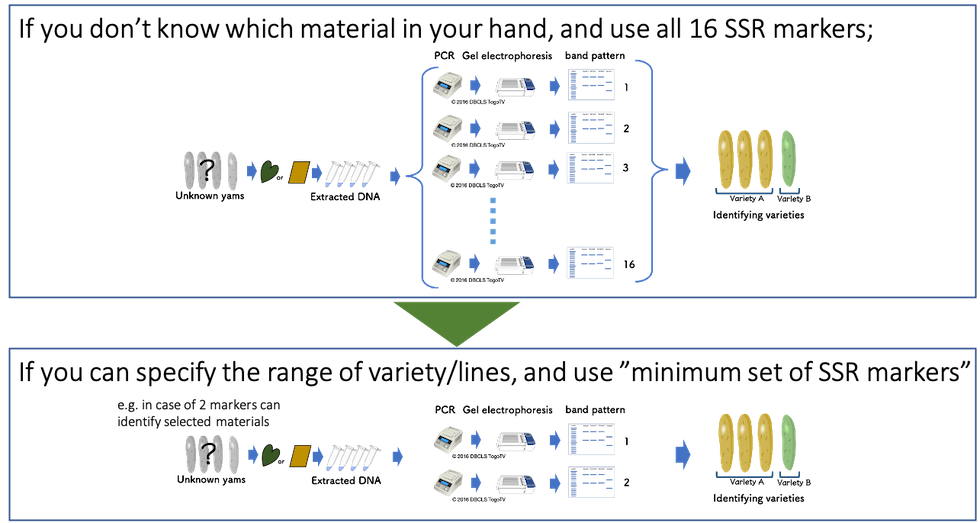
Yam Variety Identification Toolkit:Step 1 | Japan International Research Center for Agricultural Sciences | JIRCAS